 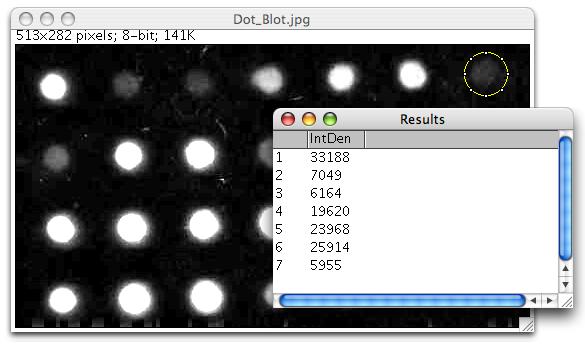
In addition, scanning software can capture the probe reactivity pattern and interpret it in terms of a genotype. Commercial instruments are available which automate the hybridization and color development. Rapid turnaround is often required in prenatal diagnosis of cystic fibrosis, beta-thalassemia and hemoglobinopathies, giving this technology has special applicability in those genetic diseases. Because of the simplicity, speed, and generally high sensitivity and specificity, large numbers of individuals can be rapidly screened using this technology. 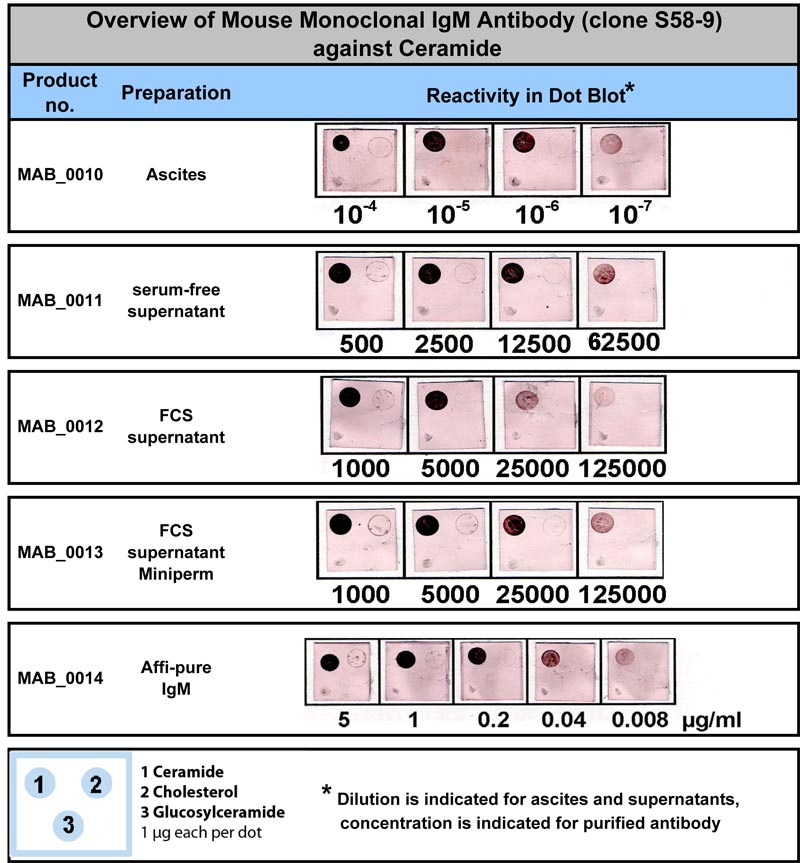
The entire procedure from blood specimen receipt to result usually requires less than 1 day. Circular spots or lines are visualized colorimetrically after hybridization through the use of streptavidin horseradish peroxidase incubation followed by development using tetramethylbenzidine and hydrogen peroxide, or via chemiluminescence after incubation with avidin alkaline phosphatase conjugate and a luminous substrate susceptible to enzyme activation, such as CSPD, followed by exposure to x-ray film. Subsequent to line or dot printing, membranes are stored or sold dry in preparation for hybridization. Sophisticated reverse dot-blot technology involves mechanized covalent attachment of activated primary amine-conjugated oligonucleotides to carboxylated nylon membranes or bovine serum albumin. In their most developed form, these assays rely upon highly multiplexed PCR reactions containing biotinylated primers providing a substrate for nonradioactive detection systems. Commercial test have been developed for human leukocyte antigens class I and class II regions of human chromosome 6, the cystic fibrosis transmembrane conductance regulator at 7q31 and strains of human Hepatitis B and C virus. After exposure, wash the membrane in TBST for 10 min.Reverse allele specific oligonucleotide assays provide a robust method for the molecular characterization of high-mutation spectrum disorders. Take the image on high-resolution mode from 1 s exposure up to 3 min exposure with appropriate intervals. Expose the membrane with your imaging system.Incubate the membrane with ECL substrate for 5 min.Wash the membrane four times for 10 min each in 10 mL of TBST with gentle shaking. The diagnostic material to be checked can be incubated on this disc. Antibody or antigen is dotted directly onto nitrocellulose membrane (NCM) discs. Flip the membrane so it is RNA-side up. Dot-immunobinding assay (Dot-Iba) is a simple and highly reproducible immunodiagnostic method.Incubate the membrane with appropriate secondary antibody (goat anti-rabbit IgG-HRP (1:20,000 dilution, ab97051) or anti-mouse IgG-HRP) in blocking buffer for 1 h at room temperature with gentle shaking. Flip the membrane so it is RNA-side down.Wash the membrane three times for 10 min each in 10 mL of TBST with gentle shaking. Flip the membrane so it is RNA-side up.Discard the blocking buffer, incubate the membrane with primary antibody (at given dilution, usually 1 μg/mL working IgG) in 10 ml blocking buffer overnight at +4☌ with gentle shaking.Incubate the membrane (RNA-side down to prevent accidental drying out of the membrane) in 10 ml blocking buffer for 1 h at room temperature with gentle shaking.Wash the membrane in 10 mL of TBST (1X TBS, 0.1% Tween-20), for 5 min at room temperature with gentle shaking to wash off the unbound RNA.Crosslink RNA to the membrane with UV light: 125 mJoule/cm2 at 254 nM. Remove the dish lid and ensure the membrane is RNA side up. Transfer the dish with membrane immediately into the chamber of SG Linker (with 254 nM bulb).Change tips after each loading, even between the same sample. Let pipetted RNA droplet diffuse onto the membrane via surface tension. NB Avoid touching the membrane with the pipette tip. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
